FAQ and Glossary¶
Why do products have clients?¶
Clients exist in tasks and products because they serve different purposes. A task client manages the connection to the database that runs your script. On the other hand, the product’s client only handles the storage of the product’s metadata.
To enable incremental runs. Ploomber has to store the source code that generated
any given product. Storing metadata in the same database that runs your code
requires a system-specific implementation. Currently, only SQLite and PostgreSQL
are supported via ploomber.products.SQLiteRelation and
ploomber.products.PostgresRelation respectively. For these two cases,
task client and product client communicate to the same system (the database).
Hence they can initialize with the same client.
For any other database, we provide two alternatives; in both cases, the
task’s client is different from the product’s client. The first alternative
is ploomber.products.GenericSQLRelation which represents a generic
table or view and saves metadata in a SQLite database; on this case, the
task’s client is the database client (e.g., Oracle, Hive, Snowflake) but
the product’s client is a SQLite client. If you don’t need the incremental
builds features, you can use ploomber.products.SQLRelation instead
which is a product with no metadata.
Which databases are supported?¶
The answer depends on the task to use. There are two types of database clients.
ploomber.clients.SQLAlchemyClient for SQLAlchemy compatible
database and ploomber.clients.DBAPIClient for the rest (the only
requirement for DBAPIClient is a driver that implements
PEP 249.
ploomber.tasks.SQLDump supports both types of clients.
ploomber.tasks.SQLScript supports both types of clients. But if you
want incremental builds, you must also configure a product client. See the section
below for details.
ploomber.tasks.SQLUpload relies on pandas.to_sql to upload a local
file to a database. Such method relies on SQLAlchemy to work. Hence it only
supports SQLAlchemyClient.
ploomber.tasks.PostgresCopyFrom is a faster alternative to
SQLUpload when using PostgreSQL. It relies on pandas.to_sql only
to create the database, but actual data upload is done using psycopg
which calls the native COPY FROM procedure.
What are incremental builds?¶
When developing pipelines, we usually make small changes and want to see how the
the final output looks like (e.g., add a feature to a model training pipeline).
Incremental builds allow us to skip redundant work by only executing tasks
whose source code has changed since the last execution. To do so, Ploomber
has to save the Product’s metadata. For ploomber.products.File, it creates
another file in the same location, for SQL products such as
ploomber.products.SQLRelation, a metadata backend is required, which
is configured using the client parameter.
How do I specify a task with a variable number of outputs?¶
You must group the outputs into a single product and declare it as a directory.
Should tasks generate products?¶
Yes. Tasks must generate at least one product; this is typically a file but can be a table or view in a database.
If you find yourself trying to write a task that generates no outputs, consider the following options:
Merge the code that does not generate outputs with upstream tasks that generate outputs.
Use the
on_finishhook to execute code after a task executes successfully (click here to learn more).
Auto reloading code in Jupyter¶
When you import a module in Python (e.g., from module import my_function),
the system caches the code and subsequent changes to my_funcion won’t take
effect even if you run the import statement again until you restar the kernel,
which is inconvenient if you are iterating on some code stored in an external
file.
To overcome such limitation, you can insert the following at the top of your
notebook, before any import statements:
# auto reload modules
%load_ext autoreload
%autoreload 2
Once executed, any updates to imported modules will take effect if you change the source code. Note that this feature has some limitations.
Parameterizing Notebooks¶
You must first parametrize the notebook by assigning the tag parameters to an
initial cell when performing a notebook task. Note that the parameters in
the parameters cell are placeholders; they indicate the parameter names that
your script or notebook takes, but they are replaced values declared in
your pipeline.yaml file at runtime. The only exception is
the upstream parameter, which contains a list of task dependencies.
Parameterizing .py files¶
For .py files, include the # %% tags=["parameters"] comment before declaring your default variables or parameters.
# %% tags=["parameters"]
upstream = None
product = None
Note that Ploomber is compatible with all .py formats supported by jupytext. Another common alternative is the light format.
The # + marker denotes the beginning of a cell, and # - marker indicates the end of the cell. Your cell should look like this:
# + tags=["parameters"]
upstream = None
product = None
# -
If you’re using another format, check out jupytext’s documentation.
Parameterizing .ipynb files in Jupyter¶
Note
This applies to JupyterLab 3.0 and higher. For more information on parameterizing notebooks in older versions, please refer to papermill docs
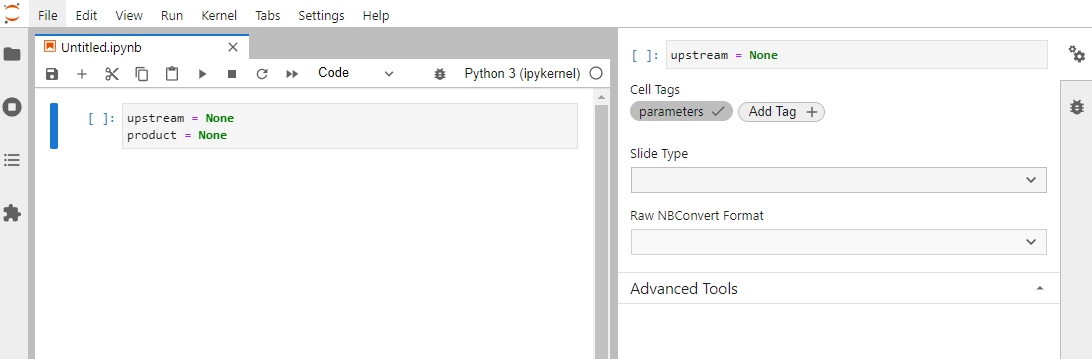
To parametrize your notebooks, add a new cell at the top, then in the right sidebar, click to open the property
inspector (double gear icon). Next, hit the “Add Tag” button, type in the word parameters, and press “Enter”.
Plotting a pipeline¶
You can generate a plot of your pipeline with ploomber plot. It supports
using D3, mermaid.js and pygraphviz as backends to create the plot. D3 is the
most straightforward option since it doesn’t require any extra dependencies, but
pygraphviz is more flexible and produces a better plot. Once installed,
Ploomber will use pygraphviz, but you can use the --backend argument
in the ploomber plot command to switch between d3, mermaid, and pygraphviz.
The simplest way to install pygraphviz is to use conda, but you can also get it working with pip.
conda (simplest)¶
conda install pygraphviz -c conda-forge
Important
If you’re running Python 3.7.x, run: conda install 'pygraphviz<1.8' -c conda-forge
pip¶
graphviz cannot be installed via pip, so you must install it with
another package manager, if you have brew, you can get it with:
brew install graphviz
Note
If you don’t have brew, refer to graphviz docs for alternatives.
Once you have graphviz, you can install pygraphviz with pip:
pip install pygraphviz
Important
If you’re running Python 3.7.x, run: pip install 'pygraphviz<1.8'
Can I use Ploomber in old JupyterLab 1.x versions?¶
Yes! Although our JupyterLab plug-in requires version 2.x, you can still
use Ploomber if using the old 1.x version, which (as of December 2021) is the
case if you’re using Amazon Sagemaker. Since Ploomber is a command-line
tool, it is independent of your editor/IDE. Furthermore, you can get the same
experience as JupyerLab users by using the
ploomber nb command; click here to learn more.
Multiprocessing errors on macOS and Windows¶
By default, Ploomber executes ploomber.tasks.PythonCallable
(i.e., function tasks) in a child process using the multiprocessing
library. On macOS and Windows, Python uses the spawn
method to create child processes; this isn’t an issue if you’re
running your pipeline from the command-line (i.e., ploomber build), but you’ll
encounter the following issue if running from a script:
An attempt has been made to start a new process before the current process
has finished its bootstrapping phase.
This probably means that you are not using fork to start your
child processes and you have forgotten to use the proper idiom
in the main module:
if __name__ == '__main__':
freeze_support()
...
The "freeze_support()" line can be omitted if the program
is not going to be frozen to produce an executable.
This happens if you store a script (say run.py):
from ploomber.spec import DAGSpec
dag = DAGSpec('pipeline.yaml').to_dag()
# This fails on macOS and Windows!
dag.build()
And call your pipeline with:
python run.py
There are two ways to solve this problem.
Solution 1: Add __name__ == '__main__'¶
To allow correct creation of child processes using spawn, run your pipeline
like this:
from ploomber.spec import DAGSpec
if __name__ == '__main__':
dag = DAGSpec('pipeline.yaml').to_dag()
# calling build under this if statement allows
# correct creation of child processes
dag.build()
Solution 2: Disable multiprocessing¶
You can disable multiprocessing in your pipeline like this:
from ploomber.spec import DAGSpec
from ploomber.executor import Serial
dag = DAGSpec('pipeline.yaml').to_dag()
# overwrite executor regardless of what the pipeline.yaml
# says in the 'executor' field
dag.executor = Serial(build_in_subprocess=False)
dag.build()
Glossary¶
Dotted path. A dot-separated string pointing to a Python module/class/function, e.g. “my_module.my_function”.
Entry point. A location to tell Ploomber how to initialize a DAG, can be a spec file, a directory, or a dotted path
Hook. A function executed after a certain event happens, e.g., the task “on finish” hook executes after the task executes successfully
Spec. A dictionary-like specification to initialize a DAG, usually provided via a YAML file